Modeling methyl-sensitive transcription factor motifs with an
Por um escritor misterioso
Descrição
Europe PMC is an archive of life sciences journal literature.

DNA methylation presents distinct binding sites for human transcription factors
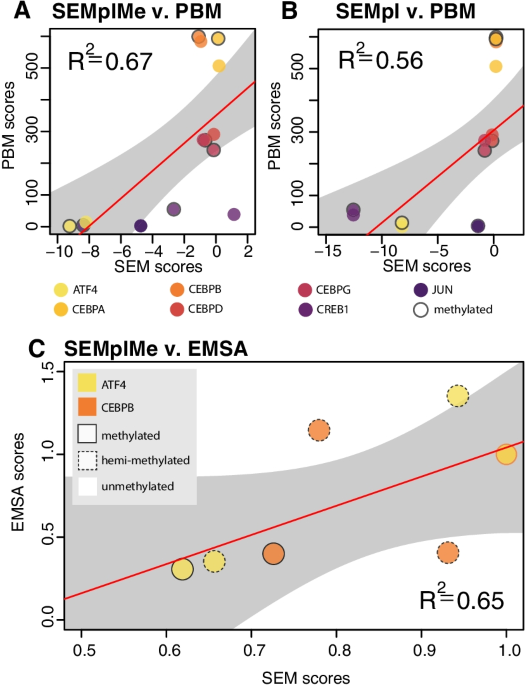
SEMplMe: a tool for integrating DNA methylation effects in transcription factor binding affinity predictions, BMC Bioinformatics

Quantitative Analysis of the DNA Methylation Sensitivity of Transcription Factor Complexes - ScienceDirect

DNA methylation: a historical perspective: Trends in Genetics

Quantitative Analysis of the DNA Methylation Sensitivity of Transcription Factor Complexes - ScienceDirect
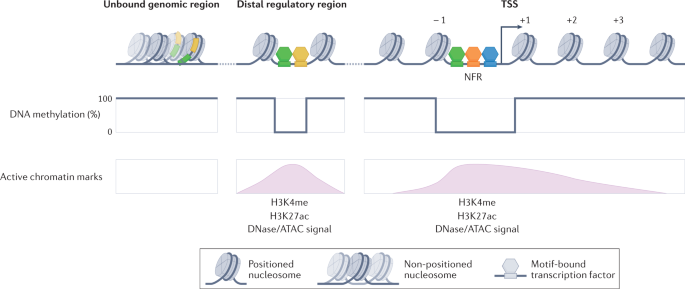
Generating specificity in genome regulation through transcription factor sensitivity to chromatin
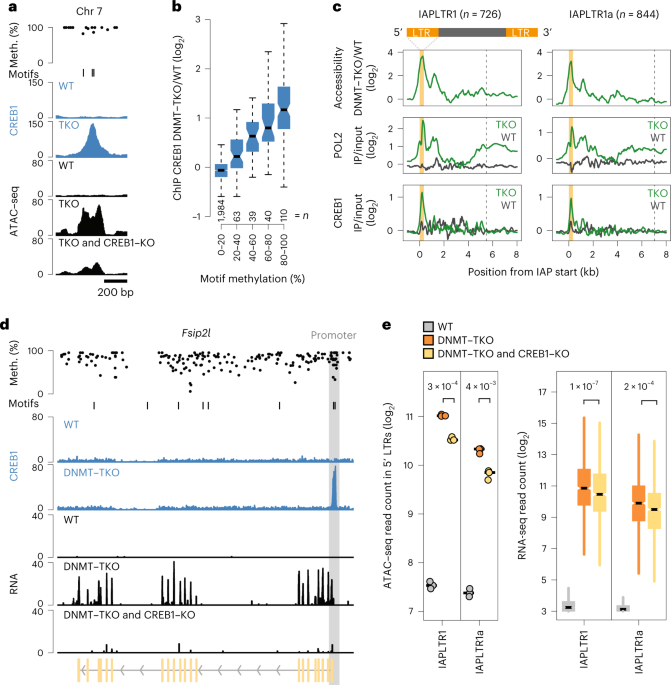
Evidence that direct inhibition of transcription factor binding is the prevailing mode of gene and repeat repression by DNA methylation
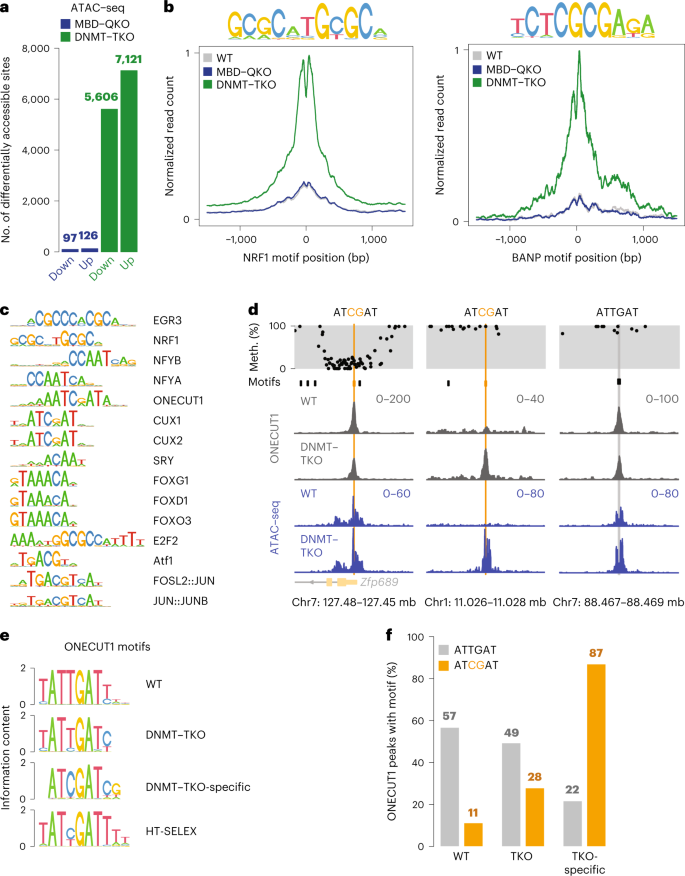
Evidence that direct inhibition of transcription factor binding is the prevailing mode of gene and repeat repression by DNA methylation
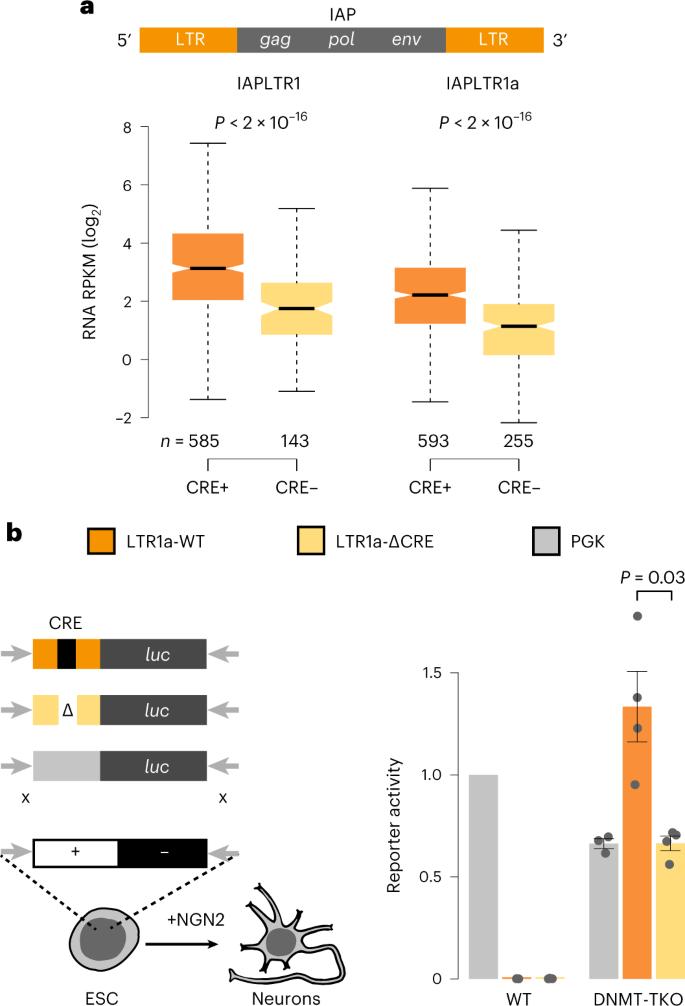
Evidence that direct inhibition of transcription factor binding is the prevailing mode of gene and repeat repression by DNA methylation

Quantitative Analysis of the DNA Methylation Sensitivity of Transcription Factor Complexes - ScienceDirect

Evidence that direct inhibition of transcription factor binding is the prevailing mode of gene and repeat repression by DNA methylation

DNA methylation patterns of transcription factor binding regions characterize their functional and evolutionary contexts
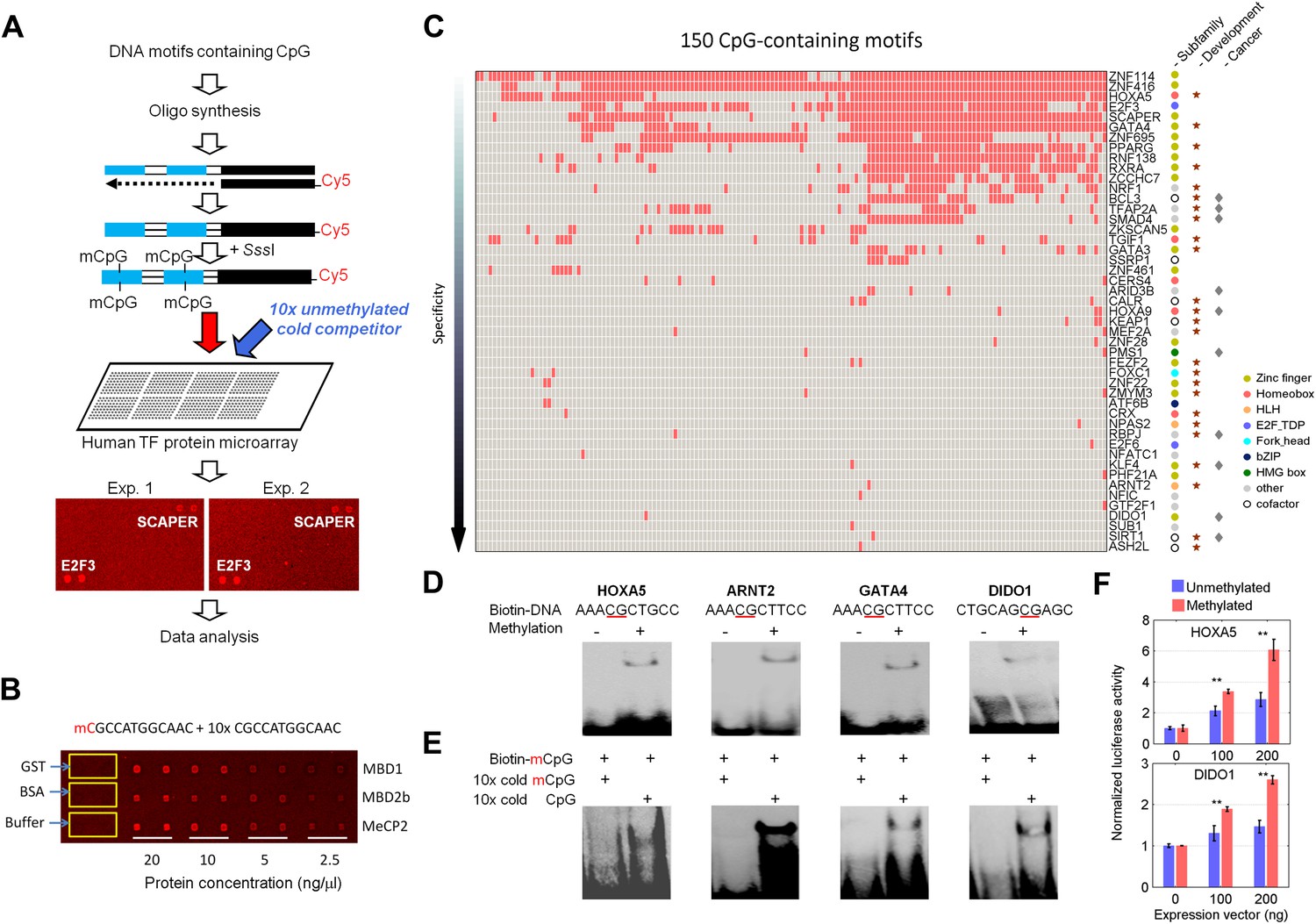
DNA methylation presents distinct binding sites for human transcription factors
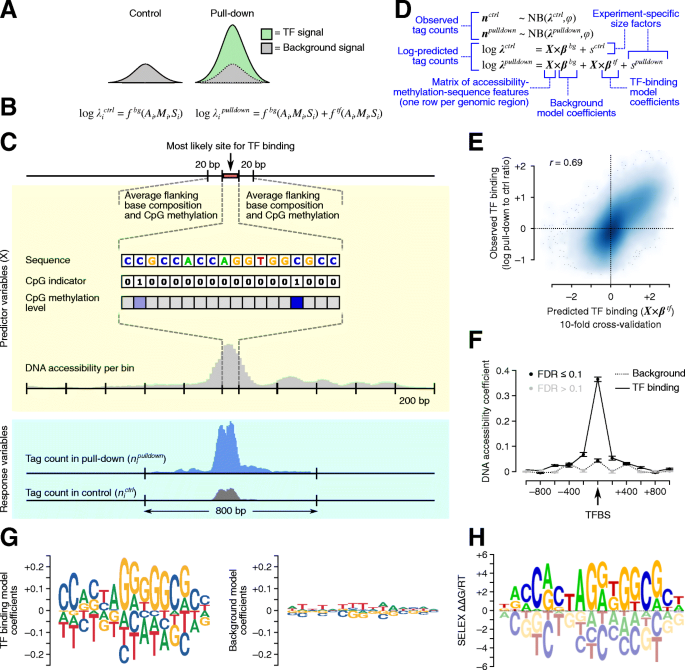
Toward a base-resolution panorama of the in vivo impact of cytosine methylation on transcription factor binding, Genome Biology

DNA methylation patterns of transcription factor binding regions characterize their functional and evolutionary contexts
de
por adulto (o preço varia de acordo com o tamanho do grupo)